Bayesian Phylogenetic Inference Explained Simply
Introduction
Bayesian phylogenetic inference is a powerful statistical method used to estimate evolutionary trees using DNA data. It calculates the probability of a tree being correct based on observed molecular sequences. Therefore, it provides a clear probability-based approach to understanding evolution. Today, this method is widely used in molecular systematics because it combines prior knowledge with statistical modeling. As a result, students find it easier to interpret compared to older tree-building methods. You can easily download this note as a PDF using the link provided just below the post for quick access and offline reading.
Evolution Notes | Systematics Notes | Evolution PPTs | Systematics PPTs
Definition
Bayesian phylogenetic inference is a statistical approach that estimates evolutionary relationships by calculating the probability of a phylogenetic tree using DNA data, evolutionary models, and prior assumptions through Bayesian probability theory.
What Is Bayesian Phylogenetic Inference?
Bayesian phylogenetic inference is based on Bayes’ Theorem. It evaluates how likely a tree is after considering the observed data.
Posterior = (Likelihood × Prior) / Evidence
- Prior represents assumptions before analyzing data.
- Likelihood shows how well a tree explains the DNA sequences.
- Posterior gives the final probability of the tree.

Unlike Maximum Parsimony, which selects the simplest tree, this method evaluates many trees. Consequently, it identifies the most probable evolutionary relationships instead of only one best tree.
Why Bayesian Phylogenetic Inference Is Important
Bayesian phylogenetic inference plays a major role in modern evolutionary studies.
- It estimates the probability of each clade.
- It applies realistic DNA evolution models.
- It provides direct statistical support values called posterior probabilities.
Two commonly used software programs are:
- MrBayes
- BEAST
These programs use advanced algorithms to analyze large molecular datasets efficiently.
You may also like NOTES in... BOTANY BIOCHEMISTRY MOL. BIOLOGY ZOOLOGY MICROBIOLOGY BIOSTATISTICS ECOLOGY IMMUNOLOGY BIOTECHNOLOGY GENETICS EMBRYOLOGY PHYSIOLOGY EVOLUTION BIOPHYSICS BIOINFORMATICS
Step-by-Step Procedure in Phylogenetic Analysis
1. Collect Molecular Data
Researchers begin by selecting DNA regions. In plant studies, common genes include:
- rbcL
- matK
- ITS region
These genes provide useful variation for evolutionary comparisons.
2. Select a Model of Evolution
Next, researchers choose an evolutionary model. This step is essential because DNA substitutions follow specific patterns.
- Jukes–Cantor model
- GTR (General Time Reversible) model
Choosing the correct model improves the reliability of results.
3. Run MCMC Analysis
The software then applies Markov Chain Monte Carlo (MCMC) analysis.
- Generates thousands or millions of possible trees
- Samples trees repeatedly
- Calculates posterior probabilities
Therefore, Bayesian phylogenetic inference does not depend on a single tree. Instead, it evaluates many possible evolutionary scenarios.
4. Interpret Posterior Probabilities
- 0.95–1.00 → Strong support
- 0.80–0.94 → Moderate support
- Below 0.80 → Weak support
These values are easier to understand than bootstrap percentages used in other methods.
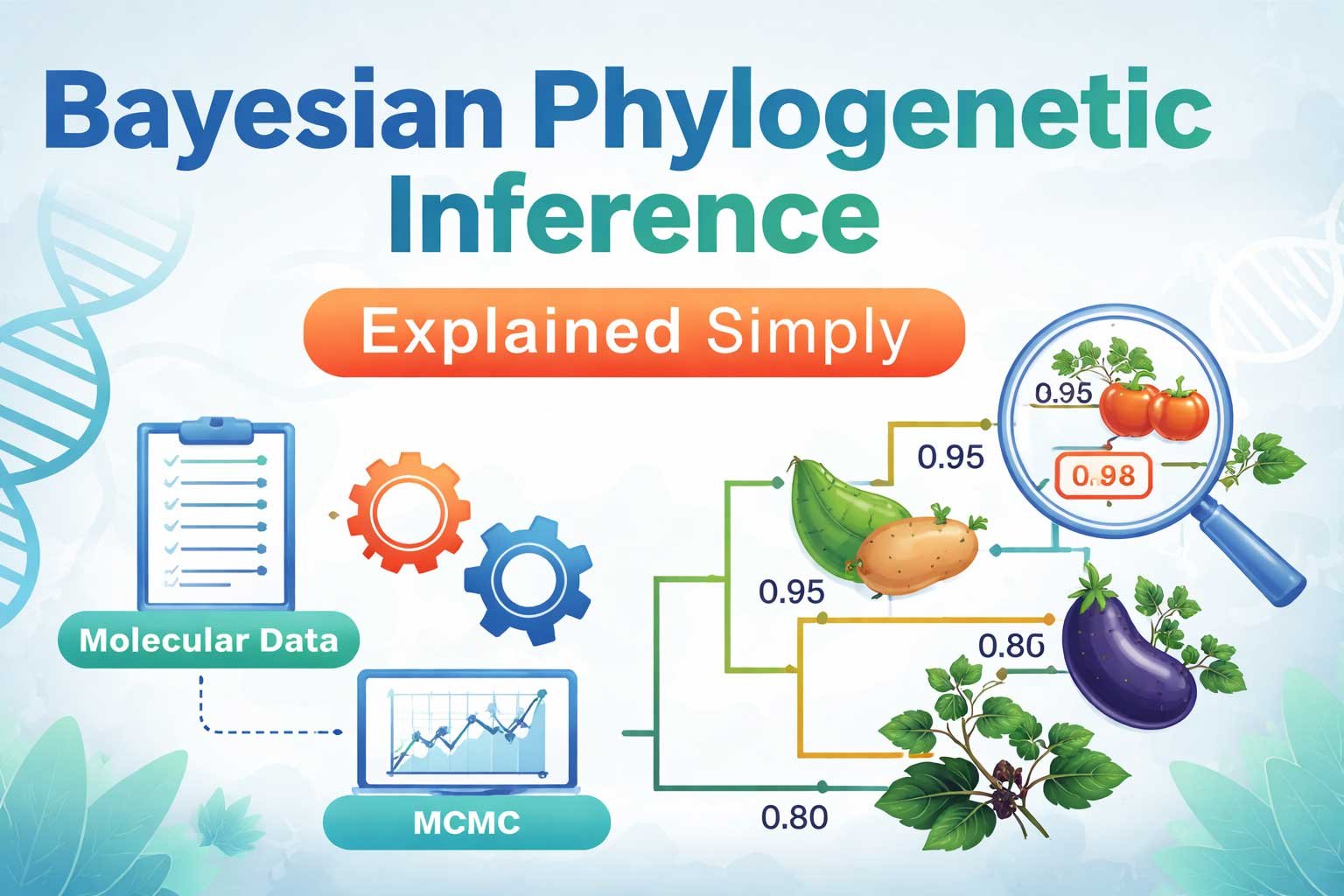
Example: Plant Taxonomy in Solanum
To understand Bayesian phylogenetic inference better, consider the genus Solanum.
- Solanum tuberosum (potato)
- Solanum melongena (brinjal)
- Solanum nigrum
- Solanum lycopersicum (tomato)
Suppose researchers analyze the rbcL gene using MrBayes.
Results May Show:
- Potato and tomato cluster together with probability 0.98
- Brinjal forms a sister group with probability 0.95
- S. nigrum appears as a separate lineage
Thus, scientists conclude there is strong evidence that potato and tomato share a recent common ancestor. Consequently, taxonomists can revise classifications if needed.
Comparison with Other Phylogenetic Methods
| Method | Basis | Output |
|---|---|---|
| Maximum Parsimony | Fewest changes | Single best tree |
| Maximum Likelihood | Highest likelihood | Best statistical tree |
| Bayesian Approach | Posterior probability | Set of probable trees |
Although Maximum Likelihood is statistically strong, Bayesian phylogenetic inference provides direct probability values. Therefore, many students prefer it for interpretation.
Advantages in Plant Systematics
- Handles chloroplast DNA datasets efficiently
- Incorporates rate variation among sites
- Supports complex evolutionary models
- Produces clear probability-based results
Limitations
- Computationally intensive
- Sensitive to model selection
- Influenced by prior assumptions
Conclusion
In summary, Bayesian phylogenetic inference is a probability-based method for constructing evolutionary trees. It combines prior assumptions with molecular data to estimate posterior probabilities. As a result, it provides statistically supported evolutionary relationships. For students of plant systematics and molecular biology, understanding Bayesian phylogenetic inference is essential for modern evolutionary analysis.
You may also like... NOTES QUESTION BANK COMPETITIVE EXAMS. PPTs UNIVERSITY EXAMS DIFFERENCE BETWEEN.. MCQs PLUS ONE BIOLOGY NEWS & JOBS MOCK TESTS PLUS TWO BIOLOGY PRACTICAL
Study Offline!! Download this Notes as a PDF
🌿 Dear Readers,
I hope you found this article helpful and easy to understand. If you have any questions, suggestions, or thoughts, I would truly love to hear from you.
Please share your feedback in the comments below. Your participation helps make EasyBiologyClass a better learning space for everyone.
Best regards,
EasyBiologyClass